| Link type | Probability | Chain A piercings | Chain B piercings | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
| view details |
|
Unlink | 56% | +37B -108B +220B -413B | +76A -282A | ||||||||
Chain A Sequence |
AASIFAAVPRAPPVAVFKLTADFREDGDSRKVNLGVGAYRTDEGQPWVLPVVRKVEQLIAGNGSLNHEYLPILGLPEFRANASRIALGDDSPAIAQKRVGSVQGLGGTGALRIGAEFLRRWYNGNNNTATPVYVSSPTWENHNSVFMDAGFKDIRTYRYWDAAKRGLDLQGLLSDMEKAPEFSIFILHACAHNPTGTDPTPDEWKQIAAVMKRRCLFPFFDSAYQGFASGNLEKDAWAVRYFVSEGFELFCAQSFSKNFGLYNERVGNLSVVGKDEDNVQRVLSQMEKIVRTTWSNPPSQGARIVATTLTSPQLFAEWKDNVKTMADRVLLMRSELRSRLESLGTPGTWNHITDQIGMFSFTGLNPKQVEYMIKEKHIYLMASGRINMCGLTTKNLDYVAKSIHEAVTKIQ |
Chain A Sequence |
AASIFAAVPRAPPVAVFKLTADFREDGDSRKVNLGVGAYRTDEGQPWVLPVVRKVEQLIAGNGSLNHEYLPILGLPEFRANASRIALGDDSPAIAQKRVGSVQGLGGTGALRIGAEFLRRWYNGNNNTATPVYVSSPTWENHNSVFMDAGFKDIRTYRYWDAAKRGLDLQGLLSDMEKAPEFSIFILHACAHNPTGTDPTPDEWKQIAAVMKRRCLFPFFDSAYQGFASGNLEKDAWAVRYFVSEGFELFCAQSFSKNFGLYNERVGNLSVVGKDEDNVQRVLSQMEKIVRTTWSNPPSQGARIVATTLTSPQLFAEWKDNVKTMADRVLLMRSELRSRLESLGTPGTWNHITDQIGMFSFTGLNPKQVEYMIKEKHIYLMASGRINMCGLTTKNLDYVAKSIHEAVTKIQ |
| sequence length | 411,411 |
| structure length | 411,411 |
| publication title |
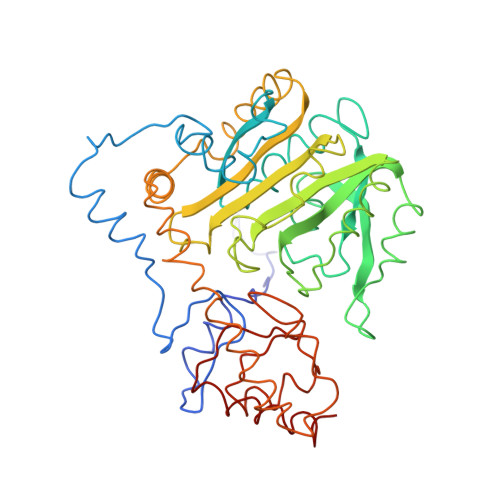
Conformational changes in cytosol aspartate aminotransferase induced by oxoglutarate
pubmed rcsb |
| molecule tags | Aminotransferase |
| molecule keywords | CYTOSOLIC ASPARTATE AMINOTRANSFERASE |
| source organism | Gallus gallus |
| ec nomenclature |
ec 2.6.1.1: Aspartate transaminase. ec 2.6.1.3: Cysteine transaminase. ec 2.6.1.1: Aspartate transaminase. ec 2.6.1.3: Cysteine transaminase. |
| pdb deposition date | 1982-04-23 |
| LinkProt deposition date | 2016-08-17 |
| chain | Pfam Accession Code | Pfam Family Identifier | Pfam Description |
|---|---|---|---|
| AB | PF00155 | Aminotran_1_2 | Aminotransferase class I and II |
| AB | PF00155 | Aminotran_1_2 | Aminotransferase class I and II |
 Image from the rcsb pdb (www.rcsb.org)
Image from the rcsb pdb (www.rcsb.org)#similar chains in the LinkProt database (?% sequence similarity) ...loading similar chains, please wait... #similar chains, but unlinked ...loading similar chains, please wait... #similar chains in the pdb database (?% sequence similarity) ...loading similar chains, please wait...