| Link type | Probability | Loop ranges | Chain A piercings | Chain B piercings | Chain C piercings | Chain D piercings | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
| view details |
|
Other | 58% | -A | +773B -852B -888D | -889A | ||||||||||
Chain A Sequence |
ETLGEKWKARLNQMSALEFYSYKKSGITEVCREEARRALKDGVATGGHAVSRGSAKLRWLVERGYLQPYGKVIDLGCGRGGWSYYAATIRKVQEVKGYTKGGPGHEEPVLVQSYGWNIVRLKSGVDVFHMAAEPCDTLLCDIGESSSSPEVEEARTLRVLSMVGDWLEKRPGAFCIKVLCPYTSTMMETLERLQRRYGGGLVRVPLSRNSTHEMYWVSGAKSNTIKSVSTTSQLLLGRMDGPRRPVKYEEDVNLGSGTRAVVSCAEAPNMKIIGNRIERIRSEHAETWFFDENHPYRTWAYHGSYEAPTQGSASSLINGVVRLLSKPWDVVTGVTGIAMTDTTPYGQQRVFKEKVDTRVPDPQEGTRQVMSMVSSWLWKELGKHKRPRVCTKEEFINKVRSNAALGAIFEEEKEWKTAVEAVNDPRFWALVDKEREHHLRGECQSCVYNMMGKREKKQGEFGKAKGSRAIWYMWLGARFLEFEALGFLNEDHWMGRENSGGGVEGLGLQRLGYVLEEMSRIPGGRMYADDTAGWDTRISRFDLENEALITNQMEKGHRALALAIIKYTYQNKVVKVLRPAEKGKTVMDIISRQDQRGSGQVVTYALNTFTNLVVQLIRNMEAEEVLEMQDLWLLRRSEKVTNWLQSNGWDRLKRMAVSGDDCVVKPIDDRFAHALRFLNDMGKVRKDTQEWKPSTGWDNWEEVPFCSHHFNKLHLKDGRSIVVPCRHQDELIGRARVSPGAGWSIRETACLAKSYAQMWQLLYFHRRDLRLMANAICSSVPVDWVPTGRTTWSIHGKGEWMTTEDMLVVWNRVWIEENDHMEDKTPVTKWTDIPYLGKREDLWCGSLIGHRPRTTWAENIKNTVNMVRRIIGDEEKYMDYLST |
Chain A Sequence |
ETLGEKWKARLNQMSALEFYSYKKSGITEVCREEARRALKDGVATGGHAVSRGSAKLRWLVERGYLQPYGKVIDLGCGRGGWSYYAATIRKVQEVKGYTKGGPGHEEPVLVQSYGWNIVRLKSGVDVFHMAAEPCDTLLCDIGESSSSPEVEEARTLRVLSMVGDWLEKRPGAFCIKVLCPYTSTMMETLERLQRRYGGGLVRVPLSRNSTHEMYWVSGAKSNTIKSVSTTSQLLLGRMDGPRRPVKYEEDVNLGSGTRAVVSCAEAPNMKIIGNRIERIRSEHAETWFFDENHPYRTWAYHGSYEAPTQGSASSLINGVVRLLSKPWDVVTGVTGIAMTDTTPYGQQRVFKEKVDTRVPDPQEGTRQVMSMVSSWLWKELGKHKRPRVCTKEEFINKVRSNAALGAIFEEEKEWKTAVEAVNDPRFWALVDKEREHHLRGECQSCVYNMMGKREKKQGEFGKAKGSRAIWYMWLGARFLEFEALGFLNEDHWMGRENSGGGVEGLGLQRLGYVLEEMSRIPGGRMYADDTAGWDTRISRFDLENEALITNQMEKGHRALALAIIKYTYQNKVVKVLRPAEKGKTVMDIISRQDQRGSGQVVTYALNTFTNLVVQLIRNMEAEEVLEMQDLWLLRRSEKVTNWLQSNGWDRLKRMAVSGDDCVVKPIDDRFAHALRFLNDMGKVRKDTQEWKPSTGWDNWEEVPFCSHHFNKLHLKDGRSIVVPCRHQDELIGRARVSPGAGWSIRETACLAKSYAQMWQLLYFHRRDLRLMANAICSSVPVDWVPTGRTTWSIHGKGEWMTTEDMLVVWNRVWIEENDHMEDKTPVTKWTDIPYLGKREDLWCGSLIGHRPRTTWAENIKNTVNMVRRIIGDEEKYMDYLS |
Chain A Sequence |
ETLGEKWKARLNQMSALEFYSYKKSGITEVCREEARRALKDGVATGGHAVSRGSAKLRWLVERGYLQPYGKVIDLGCGRGGWSYYAATIRKVQEVKGYTKGGPGHEEPVLVQSYGWNIVRLKSGVDVFHMAAEPCDTLLCDIGESSSSPEVEEARTLRVLSMVGDWLEKRPGAFCIKVLCPYTSTMMETLERLQRRYGGGLVRVPLSRNSTHEMYWVSGAKSNTIKSVSTTSQLLLGRMDGPRRPVKYEEDVNLGSGTRAVVSCAEAPNMKIIGNRIERIRSEHAETWFFDENHPYRTWAYHGSYEAPTQGSASSLINGVVRLLSKPWDVVTGVTGIAMTDTTPYGQQRVFKEKVDTRVPDPQEGTRQVMSMVSSWLWKELGKHKRPRVCTKEEFINKVRSNAALGAIFEEEKEWKTAVEAVNDPRFWALVDKEREHHLRGECQSCVYNMMGKREKKQGEFGKAKGSRAIWYMWLGARFLEFEALGFLNEDHWMGRENSGGGVEGLGLQRLGYVLEEMSRIPGGRMYADDTAGWDTRISRFDLENEALITNQMEKGHRALALAIIKYTYQNKVVKVLRPAEKGKTVMDIISRQDQRGSGQVVTYALNTFTNLVVQLIRNMEAEEVLEMQDLWLLRRSEKVTNWLQSNGWDRLKRMAVSGDDCVVKPIDDRFAHALRFLNDMGKVRKDTQEWKPSTGWDNWEEVPFCSHHFNKLHLKDGRSIVVPCRHQDELIGRARVSPGAGWSIRETACLAKSYAQMWQLLYFHRRDLRLMANAICSSVPVDWVPTGRTTWSIHGKGEWMTTEDMLVVWNRVWIEENDHMEDKTPVTKWTDIPYLGKREDLWCGSLIGHRPRTTWAENIKNTVNMVRRIIGDEEKYMDYLST |
Chain A Sequence |
ETLGEKWKARLNQMSALEFYSYKKSGITEVCREEARRALKDGVATGGHAVSRGSAKLRWLVERGYLQPYGKVIDLGCGRGGWSYYAATIRKVQEVKGYTKGGPGHEEPVLVQSYGWNIVRLKSGVDVFHMAAEPCDTLLCDIGESSSSPEVEEARTLRVLSMVGDWLEKRPGAFCIKVLCPYTSTMMETLERLQRRYGGGLVRVPLSRNSTHEMYWVSGAKSNTIKSVSTTSQLLLGRMDGPRRPVKYEEDVNLGSGTRAVVSCAEAPNMKIIGNRIERIRSEHAETWFFDENHPYRTWAYHGSYEAPTQGSASSLINGVVRLLSKPWDVVTGVTGIAMTDTTPYGQQRVFKEKVDTRVPDPQEGTRQVMSMVSSWLWKELGKHKRPRVCTKEEFINKVRSNAALGAIFEEEKEWKTAVEAVNDPRFWALVDKEREHHLRGECQSCVYNMMGKREKKQGEFGKAKGSRAIWYMWLGARFLEFEALGFLNEDHWMGRENSGGGVEGLGLQRLGYVLEEMSRIPGGRMYADDTAGWDTRISRFDLENEALITNQMEKGHRALALAIIKYTYQNKVVKVLRPAEKGKTVMDIISRQDQRGSGQVVTYALNTFTNLVVQLIRNMEAEEVLEMQDLWLLRRSEKVTNWLQSNGWDRLKRMAVSGDDCVVKPIDDRFAHALRFLNDMGKVRKDTQEWKPSTGWDNWEEVPFCSHHFNKLHLKDGRSIVVPCRHQDELIGRARVSPGAGWSIRETACLAKSYAQMWQLLYFHRRDLRLMANAICSSVPVDWVPTGRTTWSIHGKGEWMTTEDMLVVWNRVWIEENDHMEDKTPVTKWTDIPYLGKREDLWCGSLIGHRPRTTWAENIKNTVNMVRRIIGDEEKYMDYLST |
| sequence length | 883,882,883,883 |
| structure length | 883,882,883,883 |
| publication title |
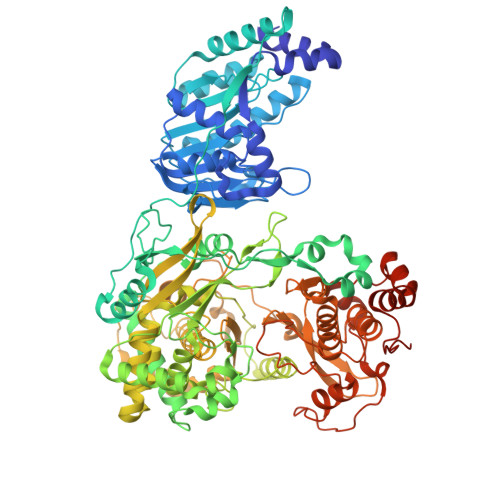
Crystal structure of the full-length Zika virus NS5 protein (Human isolate Z1106033)
rcsb |
| molecule tags | Viral protein |
| molecule keywords | NS5 |
| source organism | Zika virus (strain mr 766) |
| pdb deposition date | 2016-10-13 |
| LinkProt deposition date | 2018-06-08 |
 Image from the rcsb pdb (www.rcsb.org)
Image from the rcsb pdb (www.rcsb.org)#similar chains in the LinkProt database (?% sequence similarity) ...loading similar chains, please wait... #similar chains, but unlinked ...loading similar chains, please wait... #similar chains in the pdb database (?% sequence similarity) ...loading similar chains, please wait...